
The publications of the HotBio students
-
ESR2 Ria Desai: Deep-Sea Marine Metabolites as Promising Anti-Tubercular Agents: CADD-Guided Targeting of the F420-Dependent Oxidoreductase
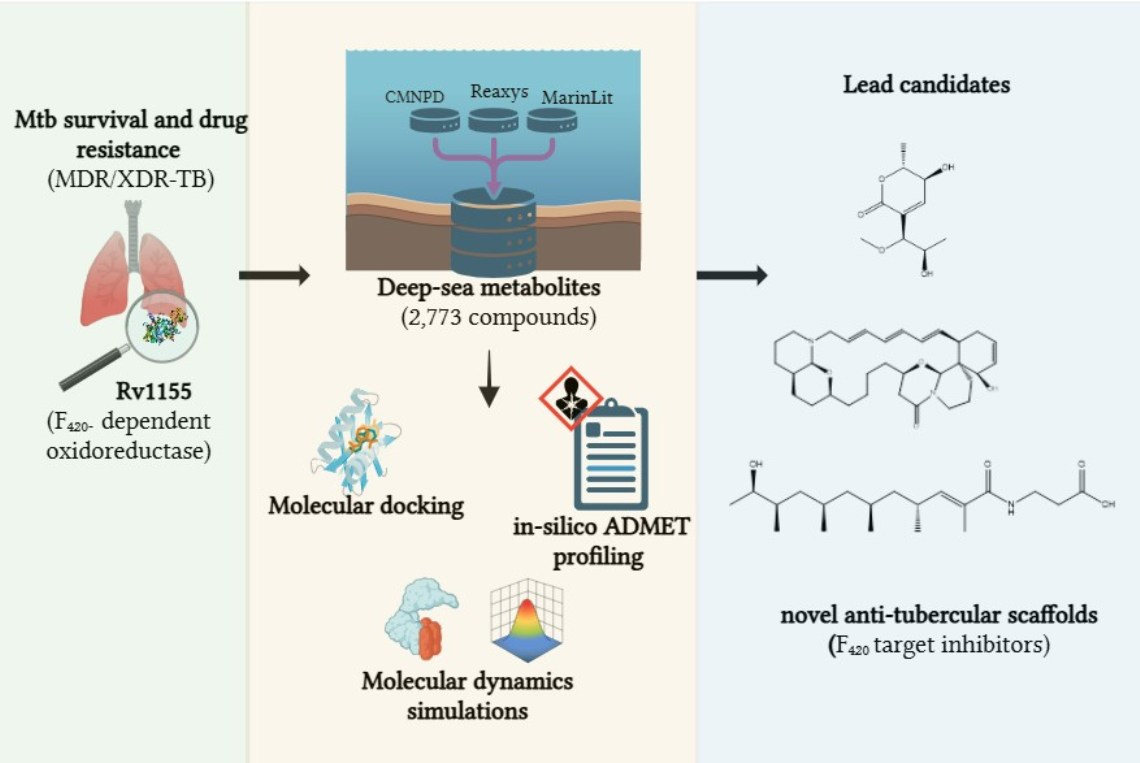
Tuberculosis (TB), caused by the bacterium Mycobacterium tuberculosis, remains a major global health challenge, especially with the rise of multidrug-resistant (MDR-TB) and extensively drug-resistant (XDR-TB) strains. To address this, Ria has been looking for new marine natural products with unique mechanisms of action. This study focuses on deep-sea marine compounds as potential inhibitors of a key enzyme in TB survival, called Rv1155.
Using advanced computer-based methods, Ria and her colleagues screened 2,773 marine-derived compounds from various chemical databases. She identified 68 compounds with strong potential to bind to Rv1155, narrowing the list down to three promising candidates: Upenamide, Aspyronol, and Fiscpropionate F. Among these, Upenamide showed the strongest and most stable binding to the enzyme, while Aspyronol demonstrated excellent drug-like properties.
This study highlights the potential of deep-sea metabolites as a source of new TB treatments and provides a cost-effective framework for further research. These findings could pave the way for experimental validation and the development of novel drugs targeting drug-resistant TB.
Desai, R.; Alaroud, A.A.; Preet, G.; Astakala, R.V.; Ebel, R.; Jaspars, M. Deep-Sea Marine Metabolites as Promising Anti-Tubercular Agents: CADD-Guided Targeting of the F420-Dependent Oxidoreductase. Mar. Drugs 2026, 24, 58. https://doi.org/10.3390/md24020058
-
ESR2 Ria Desai: An Exploratory LC-HRMS Metabolomics Study of Culture Medium-Dependent Metabolic Variation and Bioactivity in Ten Fungal Strains
Fungi represent a prolific source of structurally diverse secondary metabolites, yet the extent to which culture conditions reshape the metabolic profile and functional bioactivity remains incompletely understood.
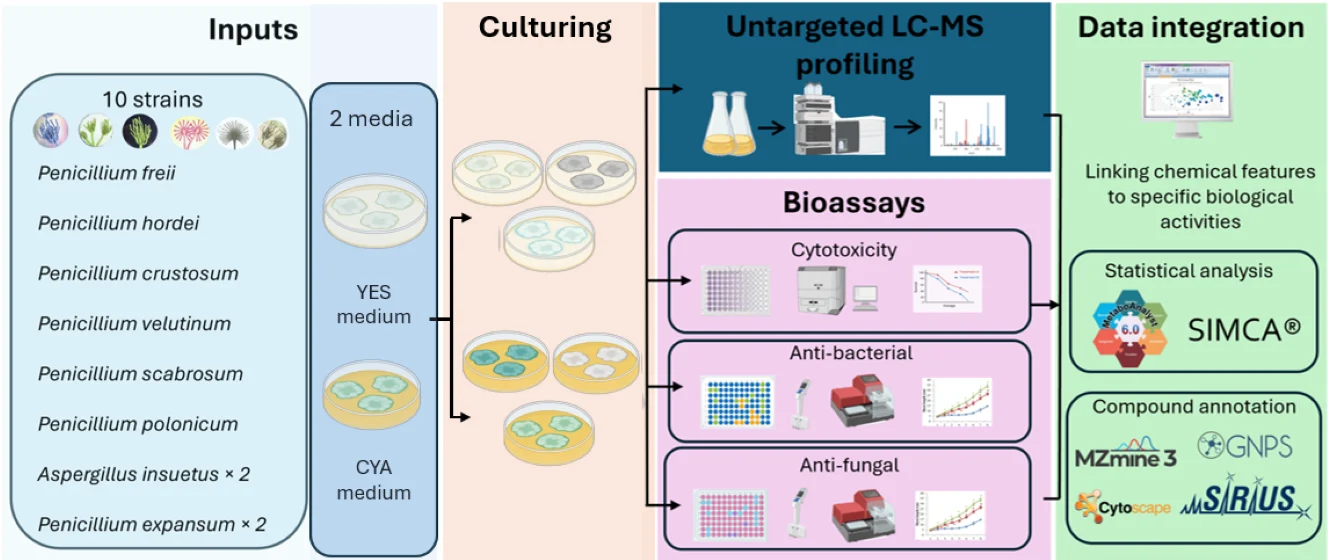
In this exploratory study, ten fungal strains belonging to genera Penicillium and Aspergillus were cultivated in Yeast Extract Sucrose (YES) and Czapek Yeast Autolysate (CYA) media and analysed using untargeted LC-HRMS metabolomics. The objective of this study was to evaluate how culture medium influences metabolic profiles and to investigate medium-dependent metabolic variation and its relation to cytotoxic, antibacterial, and antifungal activities.
Global metabolic profiling revealed moderate but statistically significant medium-associated metabolite variation, with discriminant metabolites predominantly enriched under CYA conditions. Putative structural annotation suggested patterns consistent with differential regulation of isoprenoid-derived sterols, terpenoids, alkaloid-like metabolites, and aromatic polyketides. While antimicrobial activities displayed a heterogeneous, strain-dependent pattern with limited correlation to individual metabolites, cytotoxic activity co-varied with metabolite composition in OPLS regression modelling. Sterols and terpenoid-related features emerged as major contributors to cytotoxicity. Given the absence of biological replication and the limited sample size inherent to this pilot study, all findings should be considered hypothesis-generating and interpreted within an exploratory framework.
These results suggest that nutrient composition influences biosynthetic pathway activation while functional outcomes remain strongly dependent on strain-specific metabolic capacity. This work provides a systematic framework and targeted hypothesis for future investigations into condition-dependent fungal chemical diversity in natural product discovery.
Desai, R.; Preet, G.; Astakala, R.V.; Romero-Otero, A.; Sanchez, P.; Mackenzie, T.A.; Larsen, T.O.; Ebel, R.; Jaspars, M. An Exploratory LC-HRMS Metabolomics Study of Culture Medium-Dependent Metabolic Variation and Bioactivity in Ten Fungal Strains. Int. J. Mol. Sci. 2026, 27, 3866. https://doi.org/10.3390/ijms27093866
